 B cell receptors can further diversify through somatic hypermutations during affinity maturation. V(D)J recombination selects two or three germline segments (Variable-V and Joining-J loci for TCR α and BCR light chains and the V, Diversity-D, and J loci for TCR β and BCR heavy chains) from a library of germline genes, and assembles them while deleting base pairs and inserting other non-templated ones at the junctions (Fig. Probabilistic assignment of recombination scenarios Applied to BCRs, IGoR learns a context-dependent hypermutation model to identify hotspots, which allows for a comprehensive analysis of the mutational landscape of BCRs. IGoR used as a sequence generator produces an arbitrary number of randomly rearranged sequences with the same statistics as in the data set. IGoR’s performance at identifying the correct VDJ recombination scenario is two times better than current state-of-the-art methods. Using these statistics, for each sequence IGoR outputs a whole list of potential recombination and hypermutation scenarios, with their corresponding likelihoods. We present a flexible computational method and software tool, IGoR (Inference and Generation of Repertoires), that processes raw immune sequence reads from any source (cDNA or gDNA) and learns unbiased statistics of V(D)J recombination and somatic hypermutations. Quantitatively characterizing the diversity and the biases of these mechanisms remains a challenge for understanding adaptive immunity and applying RepSeq for diagnostics. Standard assignments introduce systematic errors when describing this inherently stochastic process. Different germline segments can recombine with each other with different frequencies, and the number of insertions and deletions is random, so that the overall receptor generation process cannot be described deterministically. 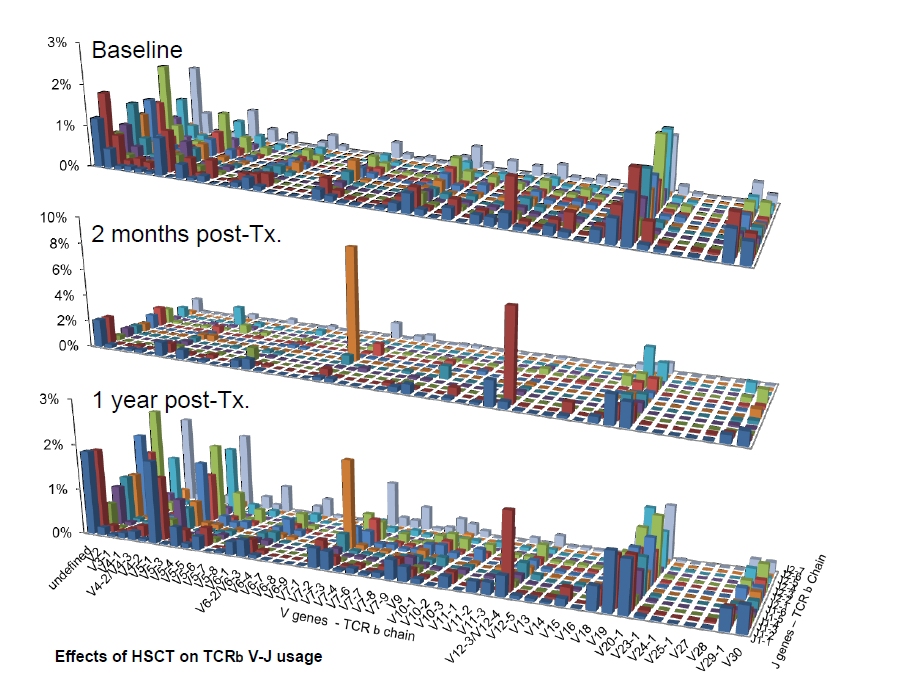  
However, each receptor sequence can be generated in a large number of ways, which we call “scenarios,” comprising the processes leading to pre-selection receptors: recombination of germline segments, insertions and deletions, and hypermutations. A growing number of algorithms and software tools have been designed to address the new challenges of RepSeq, in particular sequence analysis, germline assignment and clone construction 5, 6, 7, 8, 9, 10. The recent advent of high-throughput immune repertoire sequencing (RepSeq) 1, 2, 3, 4 gives us direct insight into the diversity of B cell and T cell receptor (BCR and TCR) repertoires with great potential to change the way we diagnose, treat, and prevent immune system-related disorders. The adaptive immune system recognizes pathogens by binding their antigens to specific surface receptors expressed on T and B cells. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
